Welcome to China Oncology,
China Oncology ›› 2024, Vol. 34 ›› Issue (4): 368-379.doi: 10.19401/j.cnki.1007-3639.2024.04.004
• Article • Previous Articles Next Articles
LIN Yicong( ), WANG Yue, XUE Qianqian, ZHENG Qiang, JIN Yan, HUANG Ziling, LI Yuan(
), WANG Yue, XUE Qianqian, ZHENG Qiang, JIN Yan, HUANG Ziling, LI Yuan( )
)
Received:2023-12-13
Revised:2024-02-25
Online:2024-04-30
Published:2024-05-17
Contact:
LI Yuan
Share article
CLC Number:
LIN Yicong, WANG Yue, XUE Qianqian, ZHENG Qiang, JIN Yan, HUANG Ziling, LI Yuan. Clinical pathological characteristics and immune microenvironment significance of EGFR T790M mutation in non-small cell lung cancer patients and its prognostic implications[J]. China Oncology, 2024, 34(4): 368-379.
Tab. 1
Clinicopathological characteristics of de novo and acquired T790M in advanced NSCLC"
| Characteristics | Total (n=82) | de novo T790M n(%) | Acquired T790M n (%) | P value |
|---|---|---|---|---|
| Total n | 82 | 15 (18.29) | 67 (81.71) | |
| Gender | 0.383 4 | |||
| Male | 32 | 4 (26.67) | 28 (41.79) | |
| Female | 50 | 11 (73.33) | 39 (58.21) | |
| Age/year | 0.255 4 | |||
| <63 | 42 | 10 (66.67) | 32 (47.76) | |
| ≥63 | 40 | 5 (33.33) | 35 (52.24) | |
| Histological differentiation | 0.026 4* | |||
| High/medium | 6 | 3 (20.00) | 3 (4.48) | |
| Low | 33 | 8 (53.33) | 25 (37.31) | |
| NA | 43 | 4 (26.67) | 39 (58.21) | |
| Specimen type | 0.036 5* | |||
| Tissue biopsy | 46 | 9 (60.00) | 37 (55.23) | |
| Surgery | 16 | 6 (40.00) | 13 (19.40) | |
| Liquid biopsy | 17 | 0 (0.00) | 17 (25.37) | |
| PD-L1 expression | 0.011 0* | |||
| Negative (<1%) | 33 | 5 (33.33) | 28 (41.79) | |
| Positive (≥1%) | 24 | 9 (60.00) | 15 (22.39) | |
| NA | 25 | 1 (6.67) | 24 (35.82) |
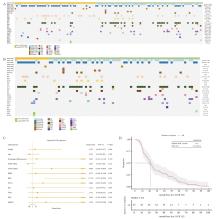
Fig. 1
Characterization of concurrent molecular alterations of de novo and acquired T790M in NSCLC A: Characterization of concurrent molecular alterations of de novo and acquired T790M in NSCLC; B: Characterization of concurrent molecular alterations of de novo and acquired T790M in advanced NSCLC; C: Forest plot of univarite Cox regression analysis in advanced NSCLC-acquired T790M; D: Median duration of remission of EGFR-TKI for acquired T790M mutation in advanced NSCLC."
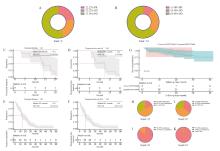


Fig. 2
The comparison between de novo and acquired T790M mutations in NSCLC concerning the utilization of third-generation EGFR TKI and subsequent prognostic analysis A: Efficacy of de novo T790M mutations in advanced NSCLC using third-generation EGFR TKI; B: Efficacy of acquired T790M mutations in advanced NSCLC using third-generation EGFR TKI; C-D: Median TD and median PFS of de novo T790M in advanced NSCLC; E-F: Median TD and median PFS of acquired T790M in advanced NSCLC; G: Survival curves for de novo and acquired T790M in advanced NSCLC; H-I: Proportion of de novo and acquired T790M recurrences; J-K: Proportion of de novo and acquired T790M metastases."

Tab. 2
Characterization of molecular alterations in advanced NSCLC acquired T790M based on efficacy groupings"
| Characteristics | Total (n = 61) | PD (n = 39) n(%) | Non-PD (n = 22) n(%) | P value |
|---|---|---|---|---|
| Total | 61 | 39 (100.00) | 22 (100.00) | |
| EGFR 19Del | 1.000 0 | |||
| Negative | 31 | 20 (51.28) | 11 (50.00) | |
| Positive | 30 | 19 (48.72) | 11 (50.00) | |
| EGFR L858R | 0.684 2 | |||
| Negative | 34 | 23 (58.97) | 11 (50.00) | |
| Positive | 27 | 16 (41.03) | 11 (50.00) | |
| TP53 | 0.528 2 | |||
| Negative | 37 | 22 (56.41) | 15 (68.18) | |
| Positive | 24 | 17 (43.59) | 7 (31.82) |
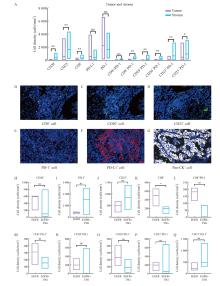
Fig. 3
Characterization of immune cell subpopulations in acquired T790M A: Comparison of immune cell density in tumor and stromal regions in acquired T790M; B-G: Schematic representation of multiple immunofluorescence histochemical expression; H-Q: Comparison of immune cell density between EGFR with tumor suppressor mutations and EGFR mutations alone."
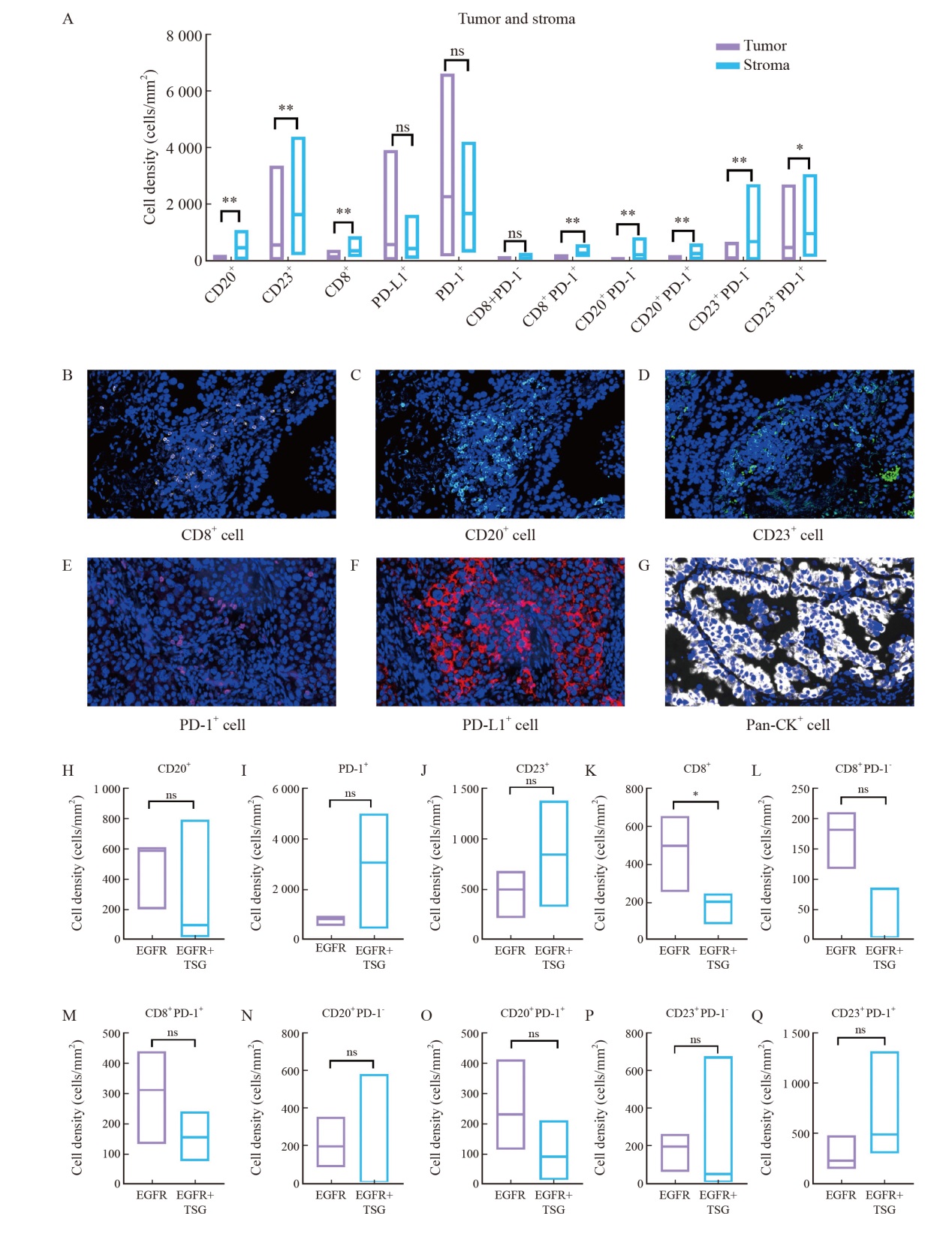
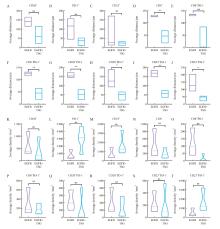
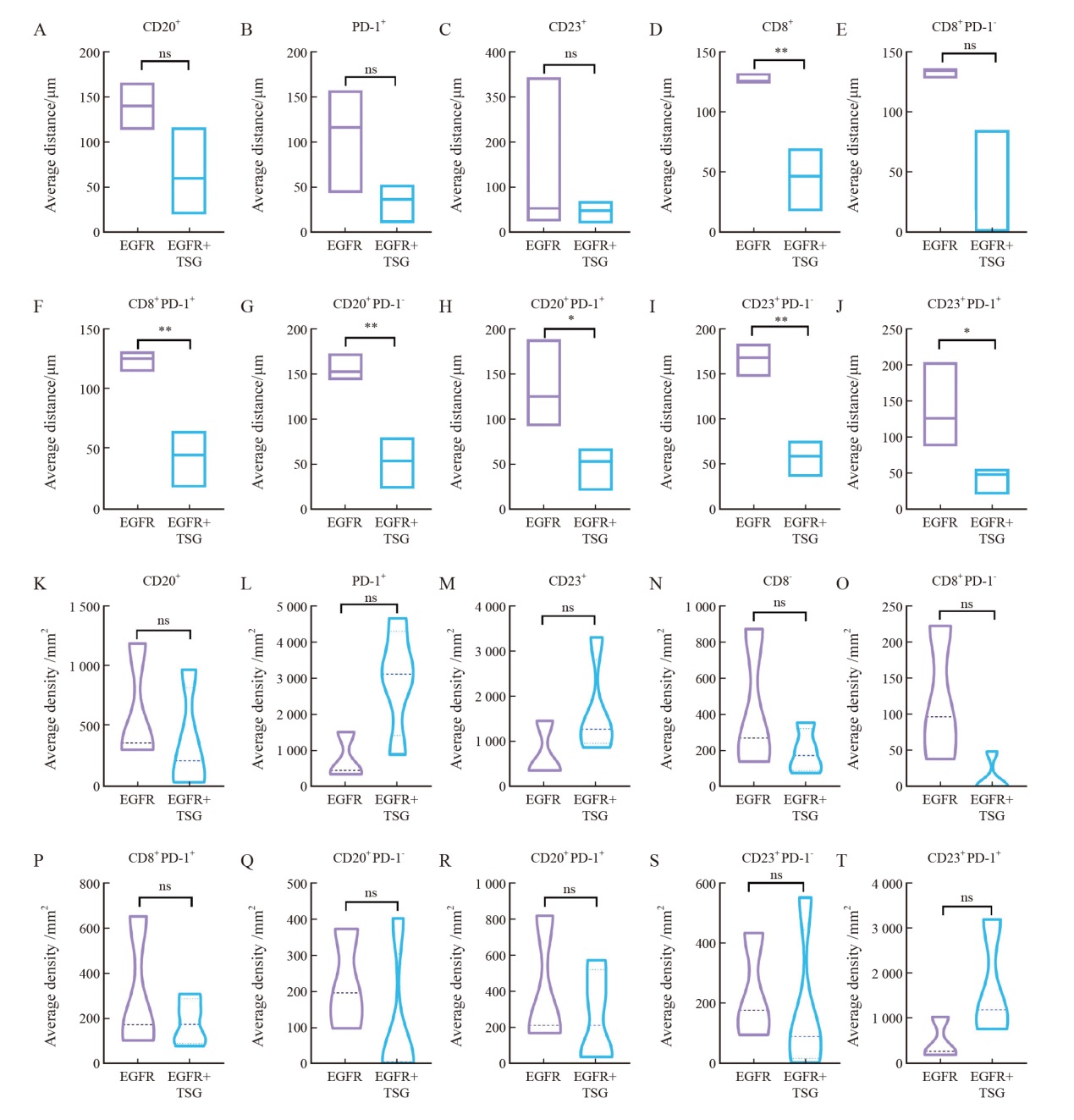
Fig. 4
Characterization of spatial distribution of immune cells in acquired T790M after treatment with EGFR TKI A-J: Comparison of the average distance between tumor cells and immune cells in 2 groups, EGFR with tumor suppressor gene mutations and EGFR mutations alone; K-T: Comparative assessment of immune cell infiltration density within a 200 μm radius at the peripheries of cancer nests between the EGFR with tumor suppressor gene mutation and EGFR mutation alone cohorts."
| [1] | SIEGEL R L, MILLER K D, FUCHS H E, et al. Cancer statistics, 2021[J]. CA A Cancer J Clin, 2021, 71(1): 7-33. |
| [2] |
DUMA N, SANTANA-DAVILA R, MOLINA J R. Non-small cell lung cancer: epidemiology, screening, diagnosis, and treatment[J]. Mayo Clin Proc, 2019, 94(8): 1623-1640.
doi: S0025-6196(19)30070-9 pmid: 31378236 |
| [3] | MILLER M, HANNA N. Advances in systemic therapy for non-small cell lung cancer[J]. BMJ, 2021, 375: n2363. |
| [4] | DA CUNHA SANTOS G, SHEPHERD F A, TSAO M S. EGFR mutations and lung cancer[J]. Annu Rev Pathol, 2011, 6: 49-69. |
| [5] |
HARRISON P T, VYSE S, HUANG P H. Rare epidermal growth factor receptor (EGFR) mutations in non-small cell lung cancer[J]. Semin Cancer Biol, 2020, 61: 167-179.
doi: S1044-579X(19)30302-5 pmid: 31562956 |
| [6] |
LIM S M, SYN N L, CHO B C, et al. Acquired resistance to EGFR targeted therapy in non-small cell lung cancer: mechanisms and therapeutic strategies[J]. Cancer Treat Rev, 2018, 65: 1-10.
doi: S0305-7372(18)30016-1 pmid: 29477930 |
| [7] | LI W H, QIU T, GUO L, et al. Primary and acquired EGFR T790M-mutant NSCLC patients identified by routine mutation testing show different characteristics but may both respond to osimertinib treatment[J]. Cancer Lett, 2018, 423: 9-15. |
| [8] |
RIZVI N A, HELLMANN M D, SNYDER A, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer[J]. Science, 2015, 348(6230): 124-128.
doi: 10.1126/science.aaa1348 pmid: 25765070 |
| [9] | ISOMOTO K, HARATANI K, HAYASHI H, et al. Impact of EGFR-TKI treatment on the tumor immune microenvironment in EGFR mutation-positive non-small cell lung cancer[J]. Clin Cancer Res, 2020, 26(8): 2037-2046. |
| [10] | GUEGUEN P, METOIKIDOU C, DUPIC T, et al. Contribution of resident and circulating precursors to tumor-infiltrating CD8+ T cell populations in lung cancer[J]. Sci Immunol, 2021, 6(55): eabd5778. |
| [11] |
PATIL N S, NABET B Y, MÜLLER S, et al. Intratumoral plasma cells predict outcomes to PD-L1 blockade in non-small cell lung cancer[J]. Cancer Cell, 2022, 40(3): 289-300.e4.
doi: 10.1016/j.ccell.2022.02.002 pmid: 35216676 |
| [12] | JIN Y, XUE Q Q, SHEN X X, et al. PD-L1 expression and comprehensive molecular profiling predict survival in non-small cell lung cancer: a real-world study of a large Chinese cohort[J]. Clin Lung Cancer, 2022, 23(1): 43-51. |
| [13] | ZHENG Q, HUANG Y, ZENG X, et al. Clinicopathological and molecular characteristics associated with PD-L1 expression in non-small cell lung cancer: a large-scale, multi-center, real-worldstudy in China[J]. J Cancer Res Clin Oncol, 2021, 147(5): 1547-1556. |
| [14] | WANG X, HUI S, TAN C, et al. Comprehensive analysis of immune subtypes reveals the prognostic value of cytotoxicity and FAP+ fibroblasts in stomach adenocarcinoma[J]. Cancer Immunol Immunother, 2023, 72(6): 1763-1778. |
| [15] | CHEN Z, FILLMORE C M, HAMMERMAN P S, et al. Non-small cell lung cancers: a heterogeneous set of diseases[J]. Nat Rev Cancer, 2014, 14(8): 535-546. |
| [16] |
GELATTI A C Z, DRILON A, SANTINI F C. Optimizing the sequencing of tyrosine kinase inhibitors (TKIs) in epidermal growth factor receptor (EGFR) mutation-positive non-small cell lung cancer (NSCLC)[J]. Lung Cancer, 2019, 137: 113-122.
doi: S0169-5002(19)30661-0 pmid: 31568888 |
| [17] | NEEL D S, BIVONA T G. Resistance is futile: overcoming resistance to targeted therapies in lung adenocarcinoma[J]. NPJ Precis Oncol, 2017, 1: 3. |
| [18] | DENIS M G, VALLÉE A, THÉOLEYRE S. EGFR T790M resistance mutation in non small-cell lung carcinoma[J]. Clin Chim Acta, 2015, 444: 81-85. |
| [19] | WANG S Y, YAN B, ZHANG Y W, et al. Different characteristics and survival in non-small cell lung cancer patients with primary and acquired EGFR T790M mutation[J]. Int J Cancer, 2019, 144(11): 2880-2886. |
| [20] | AKBAY E A, KOYAMA S, CARRETERO J, et al. Activation of the PD-1 pathway contributes to immune escape in EGFR-driven lung tumors[J]. Cancer Discov, 2013, 3(12): 1355-1363. |
| [21] | VOKES N I, CHAMBERS E, NGUYEN T, et al. Concurrent TP53 mutations facilitate resistance evolution in EGFR-mutant lung adenocarcinoma[J]. J Thorac Oncol, 2022, 17(6): 779-792. |
| [22] | GOU L Y, LI A N, YANG J J, et al. The coexistence of MET over-expression and an EGFR T790M mutation is related to acquired resistance to EGFR tyrosine kinase inhibitors in advanced non-small cell lung cancer[J]. Oncotarget, 2016, 7(32): 51311-51319. |
| [23] | ZALAQUETT Z, CATHERINE RITA HACHEM M, KASSIS Y, et al. Acquired resistance mechanisms to osimertinib: the constant battle[J]. Cancer Treat Rev, 2023, 116: 102557. |
| [24] | BRUNO T C. New predictors for immunotherapy responses sharpen our view of the tumor microenvironment[J]. Nature, 2020, 577(7791): 474-476. |
| [25] | HOU H L, QIN K, LIANG Y, et al. Concurrent TP53 mutations predict poor outcomes of EGFR-TKI treatments in Chinese patients with advanced NSCLC[J]. Cancer Manag Res, 2019, 11: 5665-5675. |
| [26] | OFFIN M, CHAN J M, TENET M, et al. Concurrent RB1 and TP53 alterations define a subset of EGFR-mutant lung cancers at risk for histologic transformation and inferior clinical outcomes[J]. J Thorac Oncol, 2019, 14(10): 1784-1793. |
| [27] | BITON J, MANSUET-LUPO A, PÉCUCHET N, et al. TP53, STK11, and EGFR mutations predict tumor immune profile and the response to anti-PD-1 in lung adenocarcinoma[J]. Clin Cancer Res, 2018, 24(22): 5710-5723. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
沪ICP备12009617
Powered by Beijing Magtech Co. Ltd
