Welcome to China Oncology,
|
||
|
Screening recurrent glioblastoma-related genes and analyzing their gene expressions in association with clinicopathological parameters and prognosis
China Oncology
2022, 32 (1):
13-23.
DOI: 10.19401/j.cnki.1007-3639.2022.01.002
Background and purpose: Glioma is the most common and malignant primary brain tumor in the central nervous system (CNS). Glioblastoma is highly malignant and aggressive, and the prognosis of patients with recurrent glioblastoma is very poor. This study aimed to screen the genes related to the recurrent glioblastoma, and analyze the relationship between their expressions, clinicopathological parameters and prognosis in glioma. Methods: By mining the relevant datasets of the primary and recurrent cases of glioblastoma in the GEO database, the differentially expressed gene (DEG) in the samples of primary and recurrent glioblastomas were screened and analyzed. All DEGs analyses were carried out in ontology function and pathway enrichment. Protein-protein interaction (PPI) network was constructed and used for screening Hub gene. Key genes were intersected by PPI network and Venn diagram, and the Gene Expression Profiling Interactive Analysis (GEPIA) and Chinese Glioma Genome Atlas (CGGA) database were analyzed for association of key gene expressions and survival status. Key genes were furtherly analyzed to determine the relationship between their expressions and clinicopathological parameters of glioma. Results: There were 40 DEG screened in the dataset GSE62153, including 34 up-regulated genes and 6 down-regulated genes. There were 19 DEG screened in the dataset GSE58399, including 16 up-regulated genes and 3 down-regulated genes. Go functional analyses showed that the DEG of GSE62153 were mainly involved in 11 physiological processes, such as central nervous system development, myelin sheath, actin binding, central nervous system myelination. The DEG of GSE58399 were mainly enriched in the positive regulation of epithelial cell migration. The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment showed that the datasets GSE62153 and GSE58399 were both enriched in histidine metabolism. By using the STRING database, the core of PPI network was constructed with 20 protein molecules. A total of 10 hub genes were screened, including MOBP, OPALIN, ERMN, PLP1, MOG, CLDN11, ASPA, TMEM125, KLK6 and NKX6-2 gene. The key genes for recurrent glioblastoma were ERMN, MOG and MOBP gene. Based on analyses using The Cancer Genome Atlas (TCGA) and CGGA databases, the prognosis of patients with high expressions of ERMN, MOG and MOBP was favorable compared with the low expression group. The expression levels of key genes in glioblastoma were lower compared with the control tissues (P<0.001). There were significant differences in the expressions of ERMN, MOG and MOBP gene among different World Health Organization (WHO) grades (WHO Ⅱ, Ⅲ and Ⅳ) (P<0.001). As the grade of glioblastoma increased, the expressions of ERMN, MOG and MOBP were decreased gradually. The expressions of ERMN, MOG and MOBP gene were correlated with WHO classification, isocitrate dehydrogenase (IDH) status and clinicopathological characteristics (P<0.001). The expression of MOBP gene was correlated with age (P<0.001) and MGMT methylation status (P=0.022). Conclusion: ERMN, MOG and MOBP gene may function as tumor suppressor genes and participate in the recurrence of glioblastoma. The histidine metabolism pathway may be related to the sensitivity of methotrexate treatment. 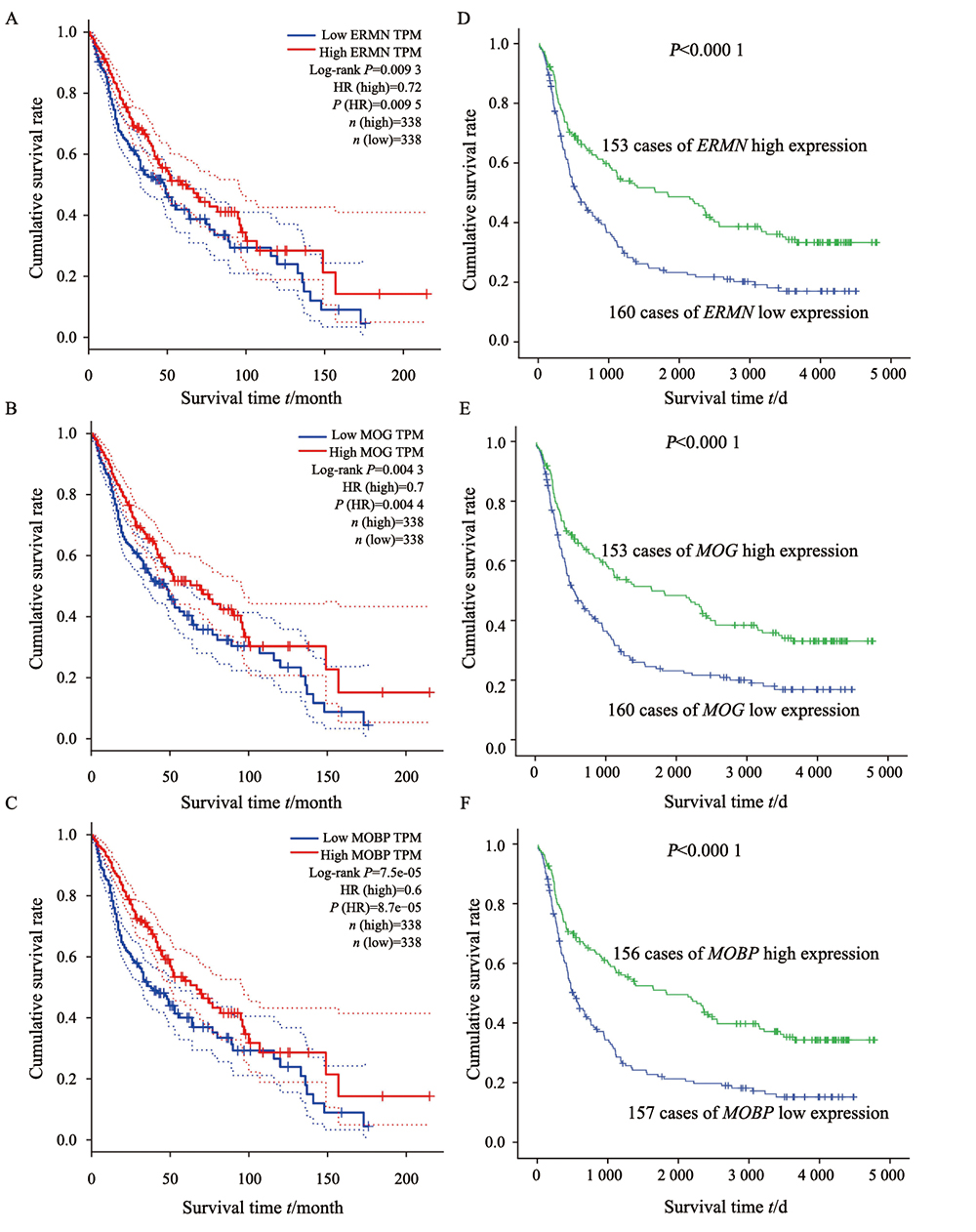
Fig. 5
Survival analysis of TCGA and CGGA database
The higher expression groups were favorable than those of patients with lower expression. A: Analysis of ERMN expression in TCGA database; B: Analysis of MOG expression in TCGA database; C: Analysis of MOBP expression in TCGA database; D: Analysis of ERMN expression in CGGA database; E: Analysis of MOG Expression in CGGA database; F: Analysis of MOBP expression in CGGA database.
Extracts from the Article
为分析ERMN、MOG和MOBP基因在胶质瘤中的预后价值,研究通过GEPIA进行生存分析,Kaplan-Meier法结果显示,在TCGA数据库676例胶质瘤病例中,ERMN、MOG和MOBP基因高表达者预后均优于低表达组,总生存率差异有统计学意义(P<0.01,图5)。为进一步验证该生存分析结果,研究进一步利用CGGA mRNA芯片数据库进行生存分析,结果显示,ERMN、MOG和MOBP基因与胶质瘤患者的总生存率均显著相关(P<0.000 1,图5),ERMN、MOG和MOBP基因高表达组预后优于低表达组。
Other Images/Table from this Article
|