Welcome to China Oncology,
China Oncology ›› 2024, Vol. 34 ›› Issue (5): 439-450.doi: 10.19401/j.cnki.1007-3639.2024.05.001
• Article • Previous Articles Next Articles
SUN Rongqi( ), SONG Ning, ZHENG Wentian, ZHANG Xinyue, LI Minmin, GONG Hui, JIANG Yingying(
), SONG Ning, ZHENG Wentian, ZHANG Xinyue, LI Minmin, GONG Hui, JIANG Yingying( )
)
Received:2023-10-07
Revised:2024-02-20
Online:2024-05-30
Published:2024-06-07
Contact:
JIANG Yingying
Share article
CLC Number:
SUN Rongqi, SONG Ning, ZHENG Wentian, ZHANG Xinyue, LI Minmin, GONG Hui, JIANG Yingying. Effect of long noncoding RNA FLJ30679 on proliferation and migration of oral squamous cell carcinoma cells[J]. China Oncology, 2024, 34(5): 439-450.
Tab. 1
The sequences of the primers used in RTFQ-PCR"
| Gene | Primer sequence |
|---|---|
| FLJ30679 | Forward: 5’-GCAAATGTGACCCGCCTCCTAC-3’ |
| Reverse: 5’-GCTACACGCTCTGCCCTTTCTC-3’ | |
| E-cadherin | Forward: 5’-CGAGAGCTACACGTTCACGG-3’ |
| Reverse: 5’-GGGTGTCGAGGGAAAAATAGG-3’ | |
| N-cadherin | Forward: 5’-TGCGGTACAGTGTAACTGGG-3’ |
| Reverse: 5’-GAAACCGGGCTATCTGCTCG-3’ | |
| Vimentin | Forward: 5’-AGTCCACTGAGTACCGGAGAC-3’ |
| Reverse: 5’-CATTTCACGCATCTGGCGTTC-3’ | |
| β-actin | Forward: 5’-CATGTACGTTGCTATCCAGGC-3’ |
| Reverse: 5’-CTCCTTAATGTCACGCACGAT-3’ | |
| GAPDH | Forward: 5’-GAACGGGAAGCTCACTGG-3’ |
| Reverse: 5’-GCCTGCTTCACCACCTTCT-3’ | |
| U6 | Forward: 5’-CTCGCTTCGGCAGCACATATACT-3’ |
| Reverse: 5’-ATTTGCGTGTCATCCTTGCGCA-3’ |
Fig. 1
Expression and prognostic significance of FLJ30679 in HNSCC tissue A: Schematic representation of TCGA data inclusion and exclusion in the UCSC Xena database. B: Analysis of FLJ30679 expression in HNSCC and normal tissues in TCGA data (Normal: 44; Cancer: 518). C: Overall survival of patients with high and low FLJ30679 expression, above the median FLJ30679 expression for the FLJ30679High group and below the median FLJ30679 expression for the FLJ30679Low group. D: Univariate COX regression analysis of FLJ30679 expression and other clinical parameters. E: Multivariate COX regression analysis of FLJ30679 expression and other clinical parameters. *: P<0.05, compared with normal. **: P<0.01, compared with normal. ****: P<0.000 1, compared with normal."
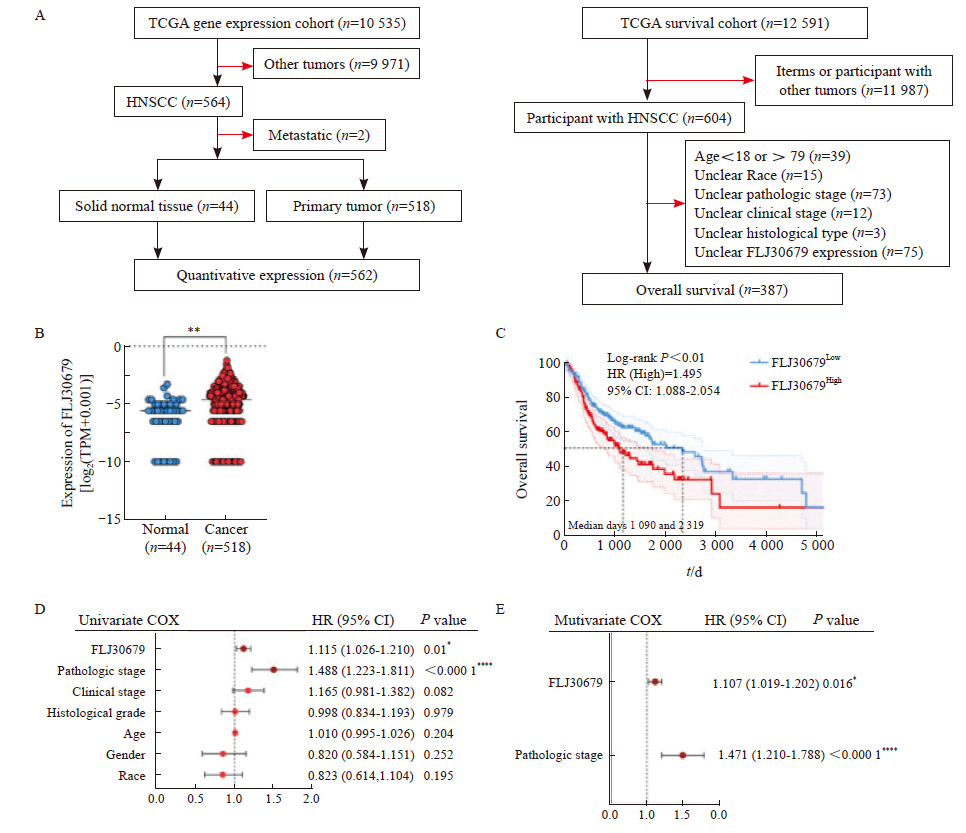

Fig. 2
Expression and subcellular localization of FLJ30679 in OSCC cell lines A: The expression of FLJ30679 in OSCC cell lines and normal cells was assessed by RTFQ-PCR. B: RNA nuclear-cytoplasmic fractionation assays were used to detect the subcellular localization of FLJ30679 in CAL-27, HN6 and HN30 cells."
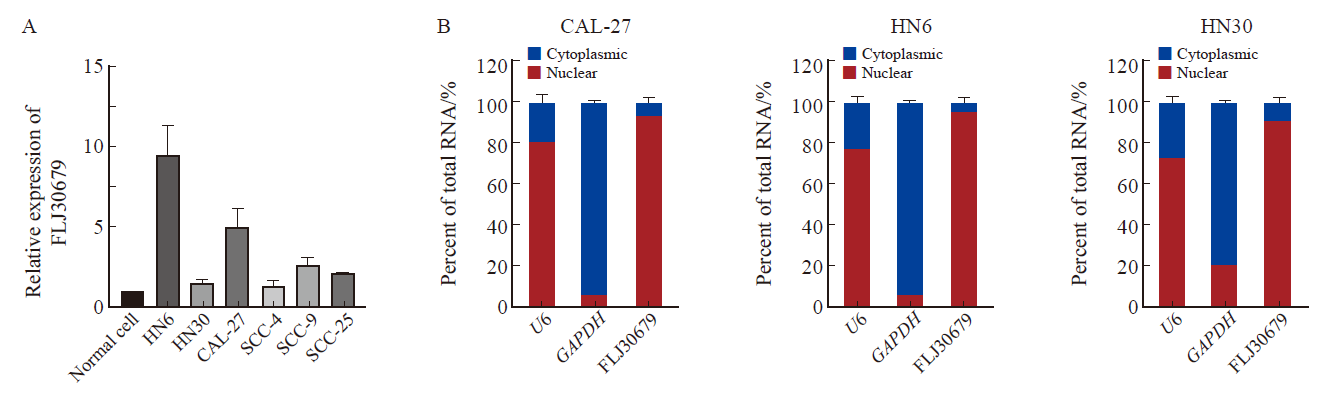
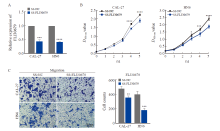
Fig. 3
Effect of FLJ30679 knockdown on proliferation and migration ability of OSCC cells A: RTFQ-PCR was used to determine the relative expression of FLJ30679 after knockdown of FLJ30679. B: CCK-8 assay was used to determine the effect of FLJ30679 knockdown on the proliferative ability of CAL-27 and HN6 cells. C: Transwell migration assay was used to determine the effect of FLJ30679 knockdown on the migratory ability of CAL-27 and HN6 cells. *: P<0.05, compared with SS-NC; **: P<0.01, compared with SS-NC; ***: P<0.001, compared with SS-NC; ****: P<0.000 1, compared with SS-NC group."
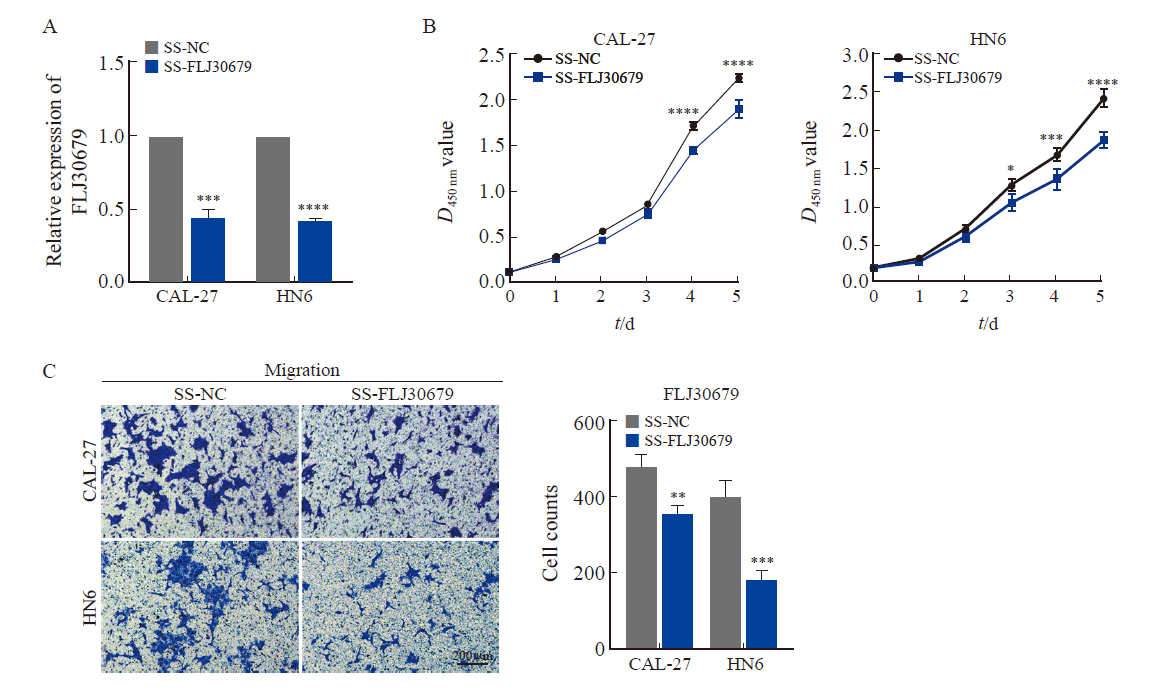
Fig. 4
Effect of FLJ30679 overexpression on proliferation and migration ability of OSCC cells A: RTFQ-PCR was used to determine the relative expression of FLJ30679 after overexpression of FLJ30679. B: CCK-8 assay was used to determine the effect of FLJ30679 overexpression on the proliferative capacity of CAL-27 and HN30 cells. C: Transwell migration assay was used to determine the effect of FLJ30679 overexpression on the migratory ability of CAL-27 and HN30 cells. **: P<0.01, compared with vector; ***: P<0.001, compared with vector; ****: P<0.000 1, compared with vector group."
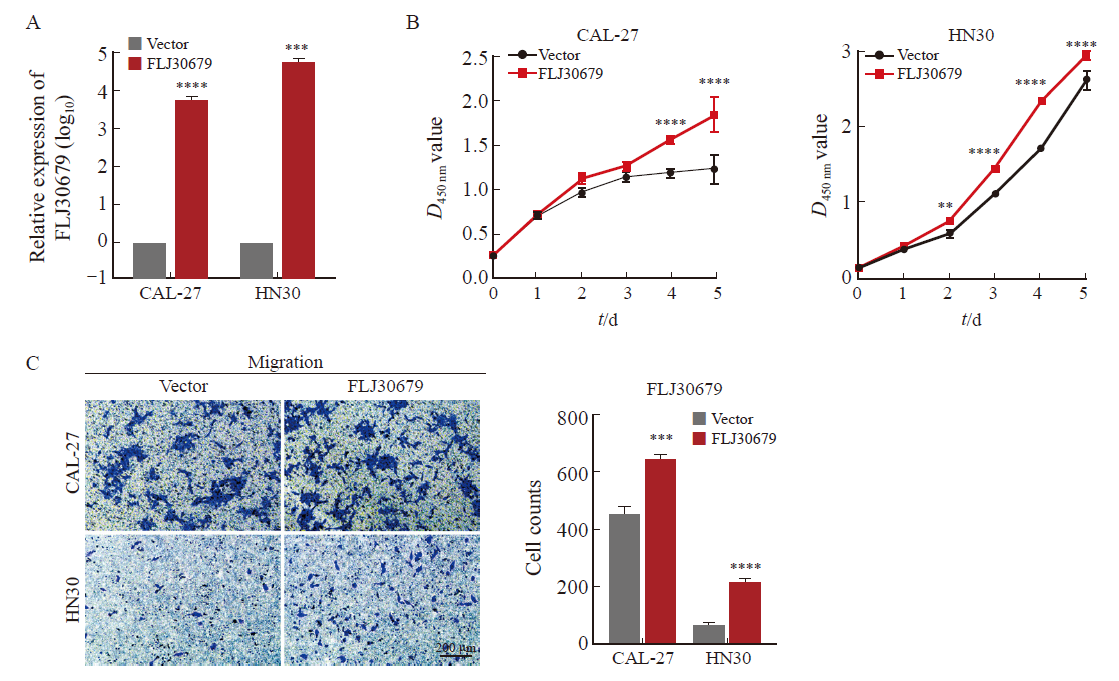
Fig. 5
Effect of FLJ30679 knockdown on the expression of EMT genes and PI3K/AKT pathway A: RTFQ-PCR was used to detect the mRNA expression levels of EMT genes after knockdown of FLJ30679. B: Western blot was used to detect the protein levels of EMT genes after knockdown of FLJ30679. C: Western blot assay used to detect the protein expression levels of PI3K/AKT and p-PI3K/p-AKT after knockdown of FLJ30679. **: P<0.01, compared with vector group or SS-NC group; ***: P<0.001, compared with vector group or SS-NC group; ****: P<0.000 1, compared with vector group or SS-NC group; NS: No significance."
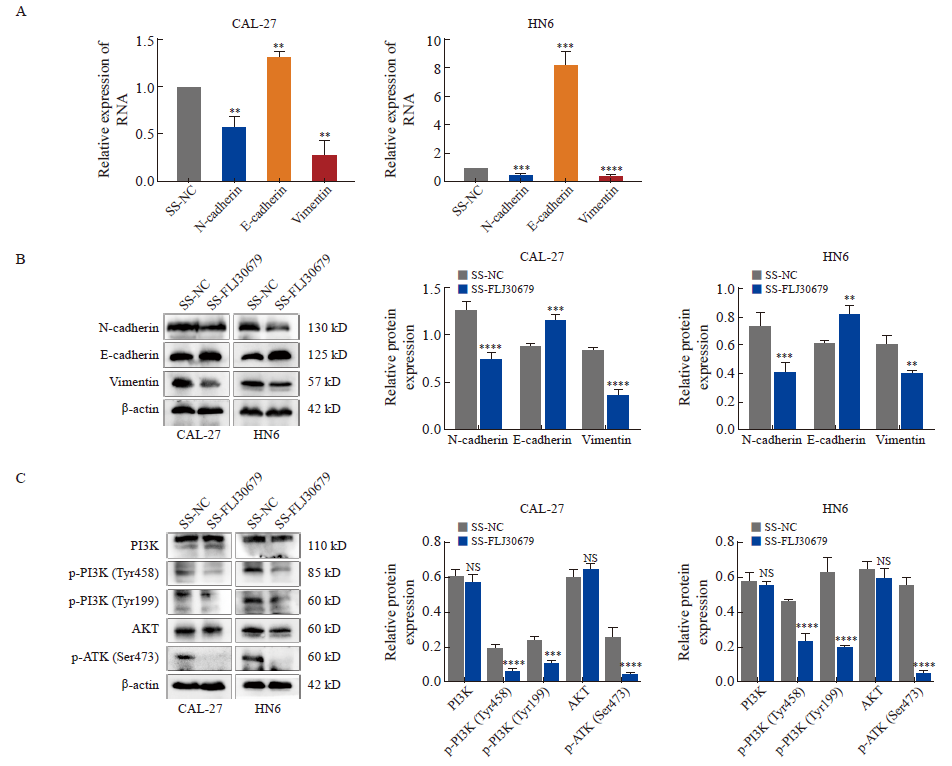
Fig. 6
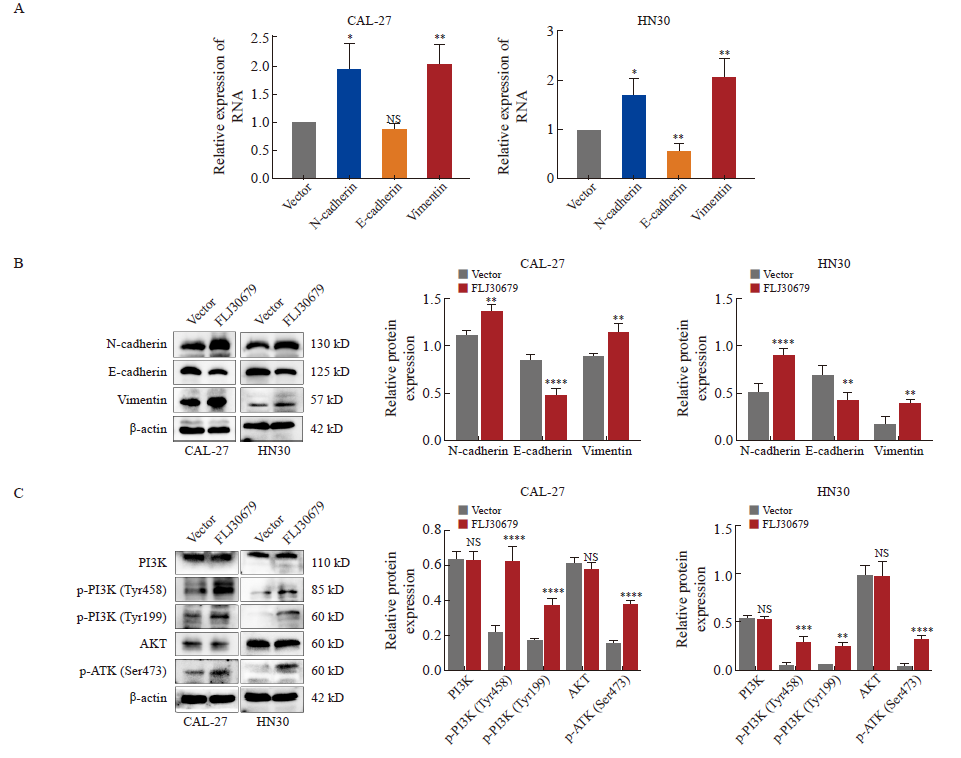
Effect of overexpression of FLJ30679 on the expression of EMT genes with PI3K/AKT pathway A: RTFQ-PCR was used to detect the mRNA expression levels of EMT genes after overexpression of FLJ30679. B: Western blot was used to detect the protein levels of EMT genes after overexpression of FLJ30679. C: Western blot was used to detect the protein expression levels of PI3K/AKT and p-PI3K/p-AKT after overexpression of FLJ30679. *: P<0.05, compared with vector group; **: P<0.01, compared with vector group; ***: P<0.001, compared with vector group; ****: P<0.000 1, compared with vector group; NS: No significance."
| [1] |
CHAI A W Y, LIM K P, CHEONG S C. Translational genomics and recent advances in oral squamous cell carcinoma[J]. Semin Cancer Biol, 2020, 61: 71-83.
doi: S1044-579X(19)30260-3 pmid: 31542510 |
| [2] | PANARESE I, AQUINO G, RONCHI A, et al. Oral and oropharyngeal squamous cell carcinoma: prognostic and predictive parameters in the etiopathogenetic route[J]. Expert Rev Anticancer Ther, 2019, 19(2): 105-119. |
| [3] |
HE J Y, YE W, KOU N, et al. MicroRNA-29b-3p suppresses oral squamous cell carcinoma cell migration and invasion via IL32/AKT signalling pathway[J]. J Cell Mol Med, 2020, 24(1): 841-849.
doi: 10.1111/jcmm.14794 pmid: 31680452 |
| [4] | KURIHARA-SHIMOMURA M, SASAHIRA T, NAKAMURA H, et al. Zinc finger AN1-type containing 4 is a novel marker for predicting metastasis and poor prognosis in oral squamous cell carcinoma[J]. J Clin Pathol, 2018, 71(5): 436-441. |
| [5] |
KOPP F, MENDELL J T. Functional classification and experimental dissection of long noncoding RNAs[J]. Cell, 2018, 172(3): 393-407.
doi: S0092-8674(18)30048-5 pmid: 29373828 |
| [6] |
ULITSKY I, BARTEL D P. lincRNAs: genomics, evolution, and mechanisms[J]. Cell, 2013, 154(1): 26-46.
doi: 10.1016/j.cell.2013.06.020 pmid: 23827673 |
| [7] | GUTTMAN M, RINN J L. Modular regulatory principles of large non-coding RNAs[J]. Nature, 2012, 482(7385): 339-346. |
| [8] |
PUDERECKI M, SZUMIŁO J, MARZEC-KOTARSKA B. Novel prognostic molecular markers in lung cancer[J]. Oncol Lett, 2020, 20(1): 9-18.
doi: 10.3892/ol.2020.11541 pmid: 32565929 |
| [9] |
CHEN M W, LIU B Q, XIAO J B, et al. A novel seven-long non-coding RNA signature predicts survival in early stage lung adenocarcinoma[J]. Oncotarget, 2017, 8(9): 14876-14886.
doi: 10.18632/oncotarget.14781 pmid: 28122330 |
| [10] |
GOLDMAN M J, CRAFT B, HASTIE M, et al. Visualizing and interpreting cancer genomics data via the Xena platform[J]. Nat Biotechnol, 2020, 38(6): 675-678.
doi: 10.1038/s41587-020-0546-8 pmid: 32444850 |
| [11] | 蒋英英, 陈曦, 石雨, 等. 长链非编码RNA COL11A1-208对口腔鳞癌细胞增殖及侵袭的影响[J]. 解放军医学杂志, 2022, 47(9): 851-862. |
| [12] | JIANG Y Y, CHEN X, SHI Y, et al. Effect of long non-coding RNA COL11A1-208 on proliferation and invasion in oral squamous cell carcinoma cells[J]. Med J Chin People’s Liberation Army, 2022, 47(9): 851-862. |
| [13] | LI J H, LIU S, ZHOU H, et al. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data[J]. Nucleic Acids Res, 2014, 42(Database issue): D92-D97. |
| [14] | TANG Z F, LI C W, KANG B X, et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses[J]. Nucleic Acids Res, 2017, 45(W1): W98-W102. |
| [15] | LI X, WANG C X, ZHANG H, et al. circFNDC3B accelerates vasculature formation and metastasis in oral squamous cell carcinoma[J]. Cancer Res, 2023, 83(9): 1459-1475. |
| [16] | GAO C F, GUO C C, LIU K, et al. LncRNA HOTAIR promotes proliferation and migration of OSCC cells via targeting miR-126[J]. Minerva Gastroenterol, 2022, 68(2): 245-247. |
| [17] |
SHI L J, YANG Y Q, LI M Y, et al. LncRNA IFITM4P promotes immune escape by up-regulating PD-L1 via dual mechanism in oral carcinogenesis[J]. Mol Ther, 2022, 30(4): 1564-1577.
doi: 10.1016/j.ymthe.2022.01.003 pmid: 35051616 |
| [18] | YANG Y X, CHEN D, LIU H, et al. Increased expression of lncRNA CASC9 promotes tumor progression by suppressing autophagy-mediated cell apoptosis via the AKT/mTOR pathway in oral squamous cell carcinoma[J]. Cell Death Dis, 2019, 10(2): 41. |
| [19] | TAO D T, ZHANG Z X, LIU X, et al. LncRNA HOTAIR promotes the invasion and metastasis of oral squamous cell carcinoma through metastasis-associated gene 2[J]. Mol Carcinog, 2020, 59(4): 353-364. |
| [20] | JIANG Y Y, WU K, CAO W, et al. Long noncoding RNA KTN1-AS1 promotes head and neck squamous cell carcinoma cell epithelial-mesenchymal transition by targeting miR-153-3p[J]. Epigenomics, 2020, 12(6): 487-505. |
| [21] | CHEN L L. Linking long noncoding RNA localization and function[J]. Trends Biochem Sci, 2016, 41(9): 761-772. |
| [22] |
PASTUSHENKO I, BLANPAIN C. EMT transition states during tumor progression and metastasis[J]. Trends Cell Biol, 2019, 29(3): 212-226.
doi: S0962-8924(18)30201-0 pmid: 30594349 |
| [23] |
贾利晴, 葛小路, 姜琳, 等. LncRNA PKD2-2-3对肺腺癌细胞增殖、克隆形成、迁移及侵袭能力的影响[J]. 中国癌症杂志, 2023, 33(8): 717-725.
doi: 10.19401/j.cnki.1007-3639.2023.08.001 |
| [24] |
JIA L Q, GE X L, JIANG L, et al. Effects of lncRNA PKD2-2-3 on cell proliferation, clone formation, migration, and invasion of lung adenocarcinoma[J]. China Oncology, 2023, 33(8): 717-725.
doi: 10.19401/j.cnki.1007-3639.2023.08.001 |
| [25] |
THIERY J P, ACLOQUE H, HUANG R Y J, et al. Epithelial-mesenchymal transitions in development and disease[J]. Cell, 2009, 139(5): 871-890.
doi: 10.1016/j.cell.2009.11.007 pmid: 19945376 |
| [26] |
LING Z H, CHENG B, TAO X A. Epithelial-to-mesenchymal transition in oral squamous cell carcinoma: challenges and opportunities[J]. Int J Cancer, 2021, 148(7): 1548-1561.
doi: 10.1002/ijc.33352 pmid: 33091960 |
| [27] | JIANG Y Y, CAO W, WU K, et al. LncRNA LINC00460 promotes EMT in head and neck squamous cell carcinoma by facilitating peroxiredoxin-1 into the nucleus[J]. J Exp Clin Cancer Res, 2019, 38(1): 365. |
| [28] | YOKOYAMA K, KAMATA N, HAYASHI E, et al. Reverse correlation of E-cadherin and snail expression in oral squamous cell carcinoma cells in vitro[J]. Oral Oncol, 2001, 37(1): 65-71. |
| [29] |
HOXHAJ G, MANNING B D. The PI3K-AKT network at the interface of oncogenic signalling and cancer metabolism[J]. Nat Rev Cancer, 2020, 20(2): 74-88.
doi: 10.1038/s41568-019-0216-7 pmid: 31686003 |
| [30] | XU W T, YANG Z, LU N H. A new role for the PI3K/Akt signaling pathway in the epithelial-mesenchymal transition[J]. Cell Adh Migr, 2015, 9(4): 317-324. |
| [31] |
YE B, JIANG L L, XU H T, et al. Expression of PI3K/AKT pathway in gastric cancer and its blockade suppresses tumor growth and metastasis[J]. Int J Immunopathol Pharmacol, 2012, 25(3): 627-636.
pmid: 23058013 |
| [32] | FAN Q C, TIAN H, WANG Y, et al. Integrin-α5 promoted the progression of oral squamous cell carcinoma and modulated PI3K/AKT signaling pathway[J]. Arch Oral Biol, 2019, 101: 85-91. |
| [33] | CHAI S L, WEN Z H, ZHANG R X, et al. CCL25/CCR9 interaction promotes the malignant behavior of salivary adenoid cystic carcinoma via the PI3K/AKT signaling pathway[J]. PeerJ, 2022, 10: e13844. |
| [34] |
LI Q, LI B, LU C L, et al. LncRNA LINC01857 promotes cell growth and diminishes apoptosis via PI3K/mTOR pathway and EMT process by regulating miR-141-3p/MAP4K4 axis in diffuse large B-cell lymphoma[J]. Cancer Gene Ther, 2021, 28(9): 1046-1057.
doi: 10.1038/s41417-020-00267-4 pmid: 33311569 |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
沪ICP备12009617
Powered by Beijing Magtech Co. Ltd
