Welcome to China Oncology,
China Oncology ›› 2023, Vol. 33 ›› Issue (1): 45-53.doi: 10.19401/j.cnki.1007-3639.2023.01.005
• Article • Previous Articles Next Articles
WANG Xiaoxiao1( ), CHEN Xi1, LI Minmin1, SONG Ning1, SUN Dongyuan1, JIANG Yingying1,2(
), CHEN Xi1, LI Minmin1, SONG Ning1, SUN Dongyuan1, JIANG Yingying1,2( )
)
Received:2022-06-01
Revised:2022-08-03
Online:2023-01-30
Published:2023-02-13
Contact:
JIANG Yingying
Share article
CLC Number:
WANG Xiaoxiao, CHEN Xi, LI Minmin, SONG Ning, SUN Dongyuan, JIANG Yingying. Effects of NOL8 on cell proliferation, migration and invasion of oral squamous cell carcinoma[J]. China Oncology, 2023, 33(1): 45-53.
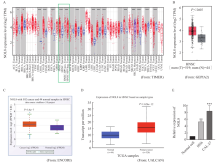
Fig. 1
The relative expression of NOL8 in OSCC tissues and cell lines A: TIMER database showed the expression of NOL8 in head and neck squamous cell carcinoma and adjacent normal tissues from green box. B: Expression of NOL8 in HNSC shown in GEPIA2 database; T: Tumor; N: Nornal. C: NOL8 expression in HNSC shown from ENCORI database. D: UALCAN database showed the expression of NOL8 in HNSC. E: RTFQ-PCR assay was used to detect the expression of NOL8 in HN6, CAL-27 and normal cell. *: P<0.05; **: P<0.01; ***: P<0.001."
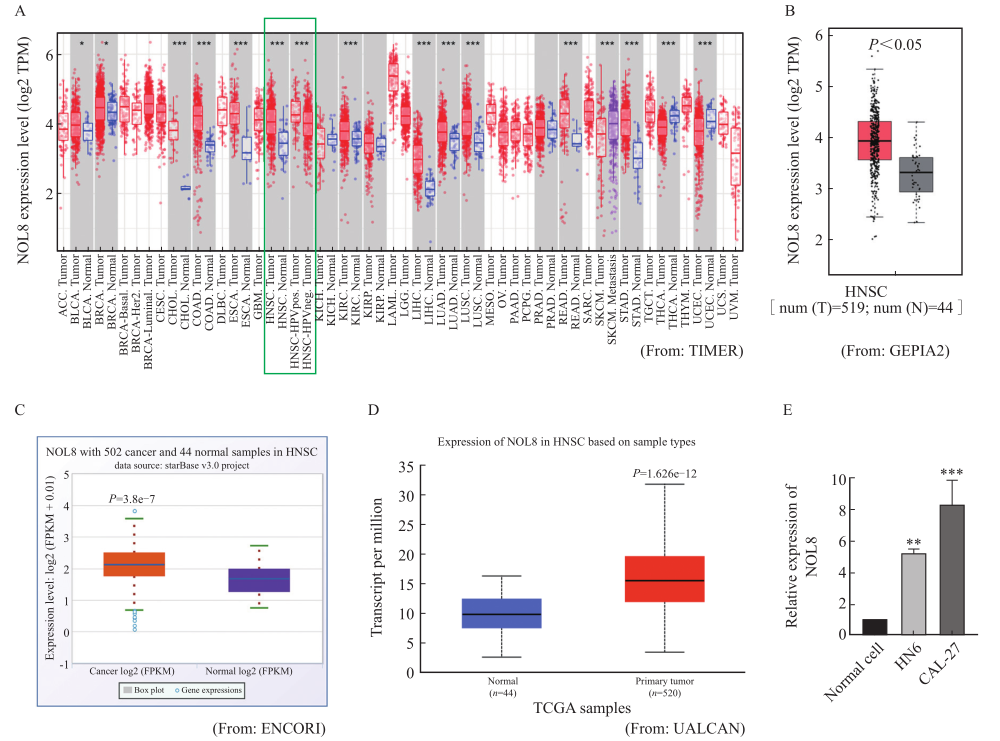
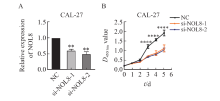
Fig. 2
Effect of knocking down NOL8 expression on the cell proliferation of OSCC A: The relative expression of NOL8 in the CAL-27 cells was detected by RTFQ-PCR after knocking down NOL8. B: CCK-8 assays were used to detect the proliferation ability of CAL-27 cells after NOL8 knockdown. **: P<0.01, compared with NC; ****: P<0.000 1, compared with si-NOL8-1 and si-NOL8-2."
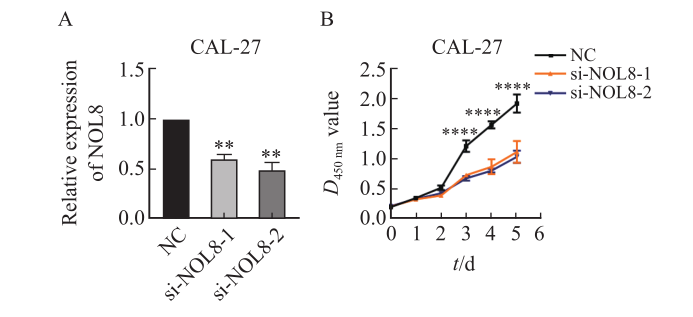
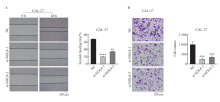
Fig. 3
Effect of knocking down NOL8 expression on cell migration and invasion of OSCC A: Scratch healing assay showed that the migration ability of CAL-27 cells after NOL8 was inhibited. B: Transwell assay showed that the invasion ability of CAL-27 cells after NOL8 was inhibited. **: P <0.01, compared with NC; ****: P<0.0001, compared with NC; ***: P<0.001, compared with NC."
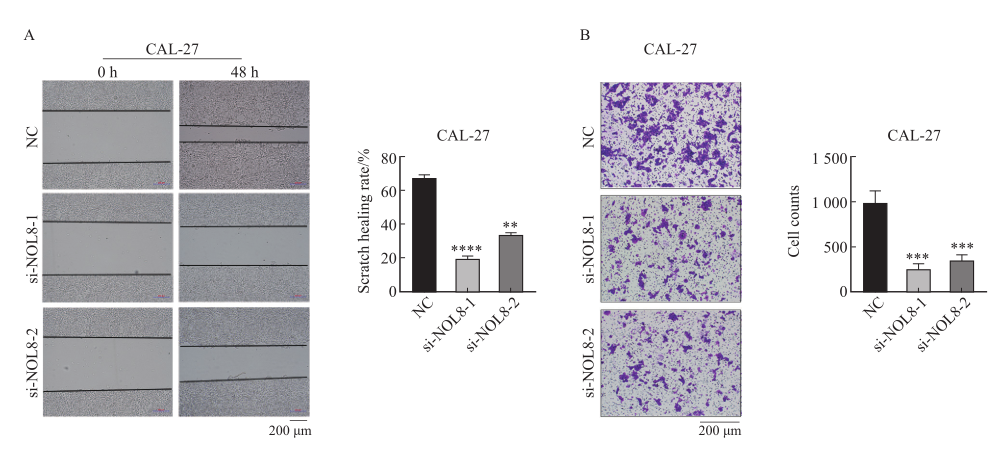

Fig. 4
Effect of overexpression of NOL8 on cell proliferation of OSCC A: The relative expression of NOL8 in the CAL-27 and HN6 cells was detected by RTFQ-PCR after over-expressing NOL8. B: CCK-8 assays were used to detect the proliferation ability of CAL-27 and HN6 cell after NOL8 was overexpressed. **: P<0.01, ****: P<0.000 1."
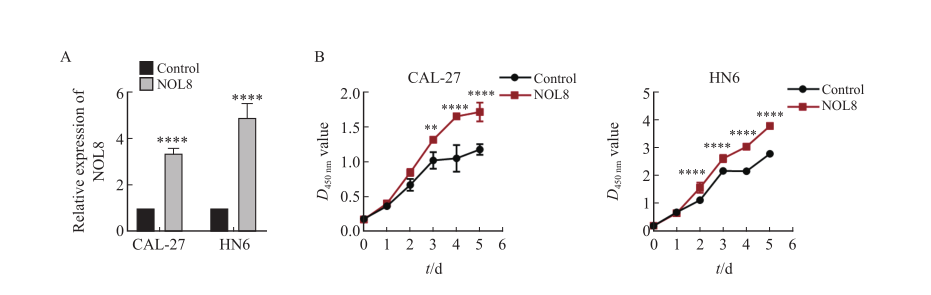
Fig. 5
Effect of overexpression of NOL8 on cell migration and invasion of OSCC A: Scratch healing assay showed that the migration ability of CAL-27 and HN6 cells after overexpression of NOL8. B: Transwell assay showed that the invasion ability of CAL-27 and HN6 cells after overexpression of NOL8. ***: P<0.001, compared with control; ****: P<0.000 1, compared with control."
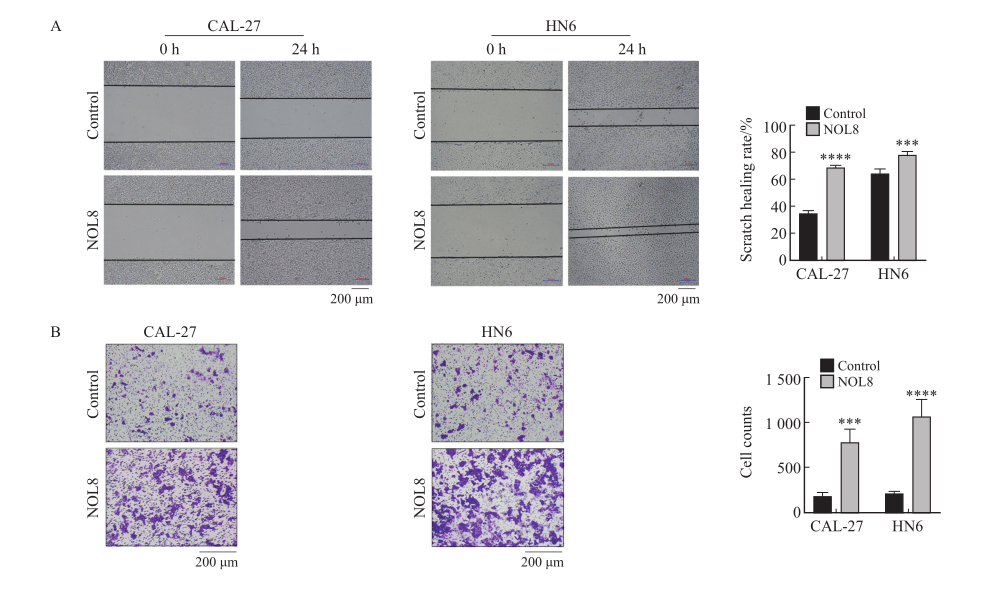
Fig. 6
Effect of the change of NOL8 expression on EMT-related proteins in CAL-27 and HN6 cells Western blot assay showed the protein level of N-cadherin, E-cadherin and vimentin after inhibition of NOL8 expression in CAL-27 cells and overexpression of NOL8 in CAL-27 and HN6 cells. **: P <0.01, compared with control; ***: P<0.001, compared with NC or control; ****: P<0.000 1, compared with NC or control."
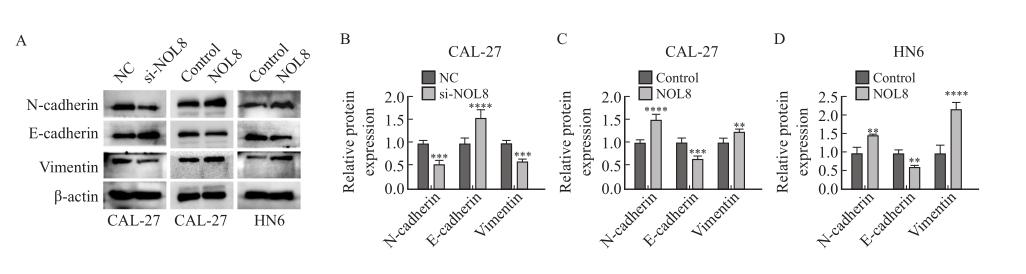
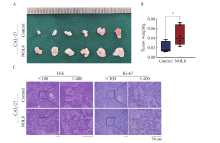
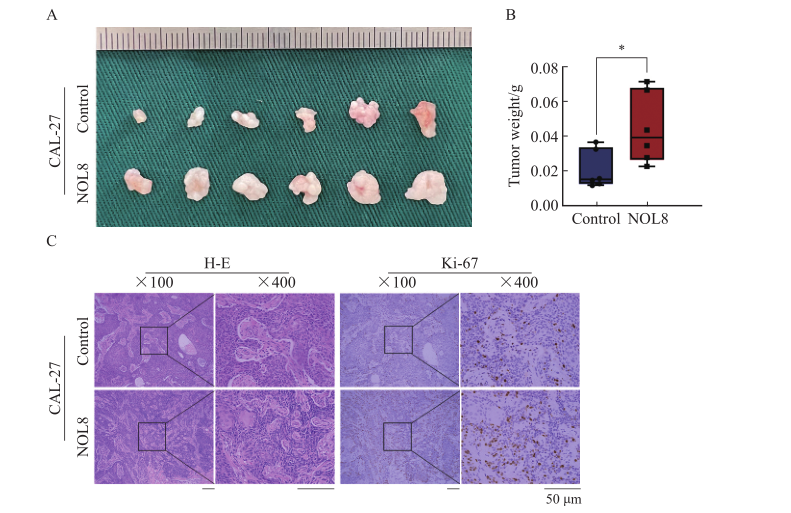
Fig. 7
Effect of NOL8 expression on the growth of subcutaneously xenograft tumor in nude mice A: OSCC xenograft tumor formed by CAL-27 cells after overexpression of NOL8. B: The weight of OSCC xenograft tumor. C: H-E and Ki-67 staining of OSCC xenograft tumor tissue of CAL-27 cells after overexpression of NOL8. *: P<0.05, compared with control."
| [1] |
CHEN W Q, ZHENG R S, BAADE P D, et al. Cancer statistics in China, 2015[J]. CA Cancer J Clin, 2016, 66(2): 115-132.
doi: 10.3322/caac.21338 |
| [2] |
AUGUSTE A, JOACHIM C, DELOUMEAUX J, et al. Head and neck cancer risk factors in the French West Indies[J]. BMC Cancer, 2021, 21(1): 1071.
doi: 10.1186/s12885-021-08787-4 pmid: 34592954 |
| [3] |
OOSTING S F, HADDAD R I. Best practice in systemic therapy for head and neck squamous cell carcinoma[J]. Front Oncol, 2019, 9: 815.
doi: 10.3389/fonc.2019.00815 pmid: 31508372 |
| [4] |
SPECENIER P, VERMORKEN J B. Optimizing treatments for recurrent or metastatic head and neck squamous cell carcinoma[J]. Expert Rev Anticancer Ther, 2018, 18(9): 901-915.
doi: 10.1080/14737140.2018.1493925 |
| [5] |
DU E, MAZUL A L, FARQUHAR D, et al. Long-term survival in head and neck cancer: impact of site, stage, smoking, and human papillomavirus status[J]. Laryngoscope, 2019, 129(11): 2506-2513.
doi: 10.1002/lary.27807 |
| [6] |
GU S, HOU P J, LIU K, et al. NOL8, the binding protein for beta-catenin, promoted the growth and migration of prostate cancer cells[J]. Chem Biol Interact, 2018, 294: 40-47.
doi: 10.1016/j.cbi.2018.08.019 |
| [7] |
MO J S, CHAE S C. microRNA 452 regulates ASB8, NOL8, and CDR2 expression in colorectal cancer cells[J]. Genes Genomics, 2021, 43(1): 33-41.
doi: 10.1007/s13258-020-01016-5 |
| [8] |
JINAWATH N, FURUKAWA Y, NAKAMURA Y. Identification of NOL8, a nucleolar protein containing an RNA recognition motif (RRM), which was overexpressed in diffuse-type gastric cancer[J]. Cancer Sci, 2004, 95(5): 430-435.
doi: 10.1111/j.1349-7006.2004.tb03227.x |
| [9] |
LU Y J, YAN Y C, LI B W, et al. A novel prognostic model for oral squamous cell carcinoma: the functions and prognostic values of RNA-binding proteins[J]. Front Oncol, 2021, 11: 592614.
doi: 10.3389/fonc.2021.592614 |
| [10] |
SEKIGUCHI T, HAYANO T, YANAGIDA M, et al. NOP132 is required for proper nucleolus localization of DEAD-box RNA helicase DDX47[J]. Nucleic Acids Res, 2006, 34(16): 4593-4608.
pmid: 16963496 |
| [11] |
O’LEARY D A, SHARIF O, ANDERSON P, et al. Identification of small molecule and genetic modulators of AON-induced dystrophin exon skipping by high-throughput screening[J]. PLoS One, 2009, 4(12): e8348.
doi: 10.1371/journal.pone.0008348 |
| [12] |
TANG Z F, LI C W, KANG B X, et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses[J]. Nucleic Acids Res, 2017, 45(W1): W98-W102.
doi: 10.1093/nar/gkx247 |
| [13] |
LI T W, FU J X, ZENG Z X, et al. TIMER2.0 for analysis of tumor-infiltrating immune cells[J]. Nucleic Acids Res, 2020, 48(W1): W509-W514.
doi: 10.1093/nar/gkaa407 |
| [14] |
CHANDRASHEKAR D S, BASHEL B, BALASUBRAMANYA S A H, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses[J]. Neoplasia, 2017, 19(8): 649-658.
doi: S1476-5586(17)30179-3 pmid: 28732212 |
| [15] |
LI J H, LIU S, ZHOU H, et al. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data[J]. Nucleic Acids Res, 2014, 42(Database issue): D92-D97.
doi: 10.1093/nar/gkt1248 |
| [16] |
HEDBERG M L, GOH G, CHIOSEA S I, et al. Genetic landscape of metastatic and recurrent head and neck squamous cell carcinoma[J]. J Clin Invest, 2016, 126(4): 1606.
doi: 10.1172/JCI86862 pmid: 27035818 |
| [17] | MASUDA K, KUWANO Y. Diverse roles of RNA-binding proteins in cancer traits and their implications in gastrointestinal cancers[J]. Wiley Interdiscip Rev RNA, 2019, 10(3): e1520. |
| [18] |
LIU J W, LI H, SHEN S X, et al. Alternative splicing events implicated in carcinogenesis and prognosis of colorectal cancer[J]. J Cancer, 2018, 9(10): 1754-1764.
doi: 10.7150/jca.24569 pmid: 29805701 |
| [19] |
MITTAL V. Epithelial mesenchymal transition in tumor metastasis[J]. Annu Rev Pathol, 2018, 13: 395-412.
doi: 10.1146/annurev-pathol-020117-043854 pmid: 29414248 |
| [20] | 周锦翰, 刘传霞, 葛巍立, 等. 上皮-间充质转化在口腔鳞状细胞癌发生、发展中作用的研究进展[J]. 浙江医学, 2021, 43(20): 2258-2262. |
| ZHOU J H, LIU C X, GE W L, et al. The role of epithelial mesenchymal transformation in the genesis and development of oral squamous cell carcinoma[J]. Zhejiang Med J, 2021, 43(20): 2258-2262. |
| [1] | WEN Ziqiang, LAN Junliang, ZHOU Bo, XU Qiwei. PARP1 promotes the progression of hepatocellular carcinoma by regulating expression of POU2F2 [J]. China Oncology, 2024, 34(9): 848-856. |
| [2] | CAO Fei, YU Wenhao, TANG Xiaonan, MA Zidong, CHANG Tingmin, GONG Yabin, LIAO Mingjuan, KANG Xiaohong. Mechanism of LINC01410 promoting proliferation and migration in esophageal squamous cell carcinoma [J]. China Oncology, 2024, 34(8): 753-762. |
| [3] | CHEN Xun, ZHENG Zhenxia, RUAN Xueru. Effects of TMCO1 on proliferation and migration of cervical cancer cells [J]. China Oncology, 2024, 34(6): 571-580. |
| [4] | SUN Rongqi, SONG Ning, ZHENG Wentian, ZHANG Xinyue, LI Minmin, GONG Hui, JIANG Yingying. Effect of long noncoding RNA FLJ30679 on proliferation and migration of oral squamous cell carcinoma cells [J]. China Oncology, 2024, 34(5): 439-450. |
| [5] | XIONG Jiayan, LEI Wei, YOU Bo, ZHANG Zhenxin, XIE Haijing, SHAN Ying, XIA Tian, ZHOU Yong. Study on the mechanism of DDX6 promoting proliferation and migration of nasopharyngeal carcinoma cells by regulating stability of CKMT1A mRNA [J]. China Oncology, 2024, 34(5): 451-459. |
| [6] | ZHOU Xueqin, LUAN Yanchao, ZHAO Li, RONG Chaochao, YANG Na. Expression of CDC20 in lung adenocarcinoma tissues and its effect on the proliferation and invasion of lung adenocarcinoma cells [J]. China Oncology, 2024, 34(5): 460-472. |
| [7] | GUAN Ruirui, HAO Qian, ZHANG Yaqi, SUN Qinggang, CHEN Yitian, LI Xiumin, ZHOU Xiang, HAN Tao. CDC20 facilitates the proliferation of esophageal carcinoma cell by stabilizing NLRP3 expression [J]. China Oncology, 2024, 34(5): 473-484. |
| [8] | WANG Xiaocong, LI Ming. The value of single-cell sequencing in oral squamous cell carcinoma research [J]. China Oncology, 2024, 34(5): 501-508. |
| [9] | XU Ziqi, HU Ruizhi, LI Junjian, WANG Hongxia, SANG Youzhou. Exploring the role of methylation-driven gene IFFO1 in pancreatic adenocarcinoma diagnosis, prognosis and cellular functions [J]. China Oncology, 2024, 34(11): 998-1010. |
| [10] | JIA Liqing, GE Xiaolu, JIANG Lin, ZHOU Xiaoyan. Effects of lncRNA PKD2-2-3 on cell proliferation, clone formation, migration, and invasion of lung adenocarcinoma [J]. China Oncology, 2023, 33(8): 717-725. |
| [11] | WANG Xuemei, CHENG Yu, QI Jiemin. PRMT7 inhibits proliferation and migration of bladder cancer cells by regulating Notch signaling pathway [J]. China Oncology, 2023, 33(5): 437-444. |
| [12] | ZHANG Pingchuan, DU Mingyu, YAO Chengyun, HE Xia, YIN Li. Mechanism of circular RNA hsa_circ_0012779 expression in nasopharyngeal carcinoma and its influence on cell biological behavior [J]. China Oncology, 2023, 33(5): 445-451. |
| [13] | XIAO Lanshu, PAN Liudi, LIU Yi, WANG Jie, CHEN Hui. LncRNA DLEU7-AS1 contributes to proliferation and migration of gastric cancer by regulating MSN transcription [J]. China Oncology, 2023, 33(4): 327-341. |
| [14] | CHEN Hong, CHEN Junxia. Effect of hsa_circ_0001573 on biological behaviors of breast cancer cells and its molecular mechanism [J]. China Oncology, 2023, 33(4): 342-353. |
| [15] | PENG Jin, WANG Weining, TAN Zhi, YE Guannan, ZHOU Zhen. The mechanism of m6Am-modifying enzyme PCIF1 regulating target gene ACOT8 in gastric cancer progression [J]. China Oncology, 2023, 33(4): 368-376. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
沪ICP备12009617
Powered by Beijing Magtech Co. Ltd
